This website was constructed as a class project for Genetics 564, an undergraduate Genetics course at UW-Madison.
Specific Aims
On this page, I describe HFI and outline projects for further understanding of the disorder as well as research for a possible HFI supplement. The Specific Aims document I submitted for Genetics 564 can be found here. The Powerpoint presentation about HFI and my Specific Aims can be found here.
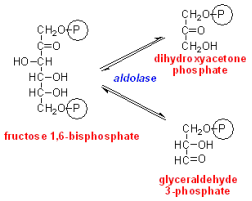
Hereditary Fructose Intolerance (HFI) is a metabolic disorder that causes affected individuals to improperly process the simple sugar, fructose [1]. The cause of this hereditary disorder is a mutation to the aldolase B gene. Aldolase B is a liver protein involved in glycolysis, the body’s process for breaking down sugars to make energy in the form of ATP [2,3]. It is important for cleaving fructose-1,6-bisphosphate, an intermediate in this process, into other intermediates, Glyceraldehyde-3-phosphate (G3P) and Dihydroxyacetone phosphate (DHAP) (see diagram at right) [1-3]. When aldolase B cannot do this, fructose builds up in the liver, causing the patient to experience nausea, vomiting, hypoglycemia, and liver damage [1,2]. HFI affects approximately 1 in 23,000 people worldwide and there is no cure or treatment available [2]. HFI patients can avoid symptoms through dietary restriction, but this can be frustrating and inconvenient.
Symptoms of disorders such as lactose intolerance can be alleviated through supplementation [4]. No such supplements are available for HFI, but a substitute for dysfunctional aldolase B could alleviate symptoms induced by fructose consumption. I hypothesize that by identifying chemicals that mimic aldolase B and the genes these chemicals affect, progress toward an HFI supplement could be made. Additionally, I hypothesize that by researching the role of specific portions of homolog aldolase B, more insight on human aldolase B could be gained.
The primary goal of this study is to identify drugs that mimic aldolase B and the genes they affect. The secondary goal of this study is to determine the importance of certain segments of the aldolase B gene.
The primary goal of this study is to identify drugs that mimic aldolase B and the genes they affect. The secondary goal of this study is to determine the importance of certain segments of the aldolase B gene.

Specific Aim 1: To identify chemicals that mimic aldolase B using a chemical genetic screen with a focused library in Drosophila.
Reason: HFI is not life-threatening when dietary modifications are made, but these modifications can sometimes be inconvenient. Identifying a chemical that essentially replaces the function of aldolase B in the body could lead to the development of a dietary supplement for individuals with HFI. This supplement would allow these individuals to consume fructose without experiencing any unpleasant symptoms associated with HFI.
Approach:
Step 1: Use morpholinos to repress the aldolase β protein in Drosophila. Morpholinos are nucleotide sequences that attach to mRNA to prevent it from being translated into a protein (see image at left). This is a useful technique because it can knock out protein function without interfering with DNA. It is an important technique for this experiment because Drosophila have only one aldolase gene for all three isozymes (α, β, and γ, homologs to human aldolase A, B, and C) instead of three separate genes, like humans.
Reason: HFI is not life-threatening when dietary modifications are made, but these modifications can sometimes be inconvenient. Identifying a chemical that essentially replaces the function of aldolase B in the body could lead to the development of a dietary supplement for individuals with HFI. This supplement would allow these individuals to consume fructose without experiencing any unpleasant symptoms associated with HFI.
Approach:
Step 1: Use morpholinos to repress the aldolase β protein in Drosophila. Morpholinos are nucleotide sequences that attach to mRNA to prevent it from being translated into a protein (see image at left). This is a useful technique because it can knock out protein function without interfering with DNA. It is an important technique for this experiment because Drosophila have only one aldolase gene for all three isozymes (α, β, and γ, homologs to human aldolase A, B, and C) instead of three separate genes, like humans.
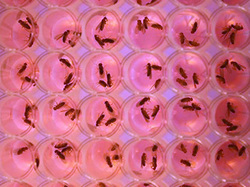
Step 2: Distribute wild-type (phenotypically normal) and aldolase β protein knockout Drosophila (created in step 1) evenly into well plates (shown below). Each well will have a different chemical from a focused library. A focused chemical library is a list of chemicals that are very similar, but vary according to one specification. The function and structure of aldolase B have been identified, and in general chemicals that are more alike are more likely to have the same function. For this experiment, it makes sense to use a focused chemical library to screen for a chemical that can function similarly to aldolase B.
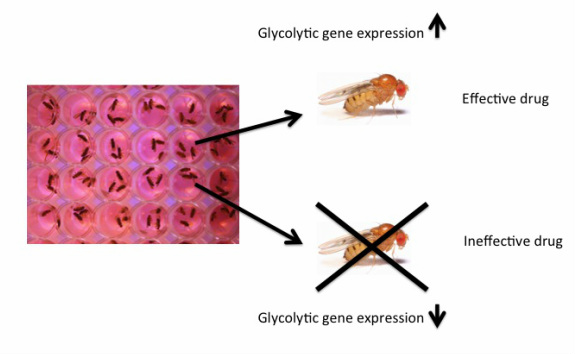
Step 3: Feed flies fructose and examine their overall phenotype and gene expression. Knockout flies that are unable to process fructose will die because the chemical they were given was ineffective at replacing the function of aldolase β in their body. Knockout Drosophila that do survive will be able to process fructose like the wild-type control Drosophila, which indicates the chemical they were given is effective at rescuing fructose metabolism. In order to determine that this chemical really is working as it should be, gene expression should be evaluated and compared to wild-type flies. It is counterproductive to fix one problem while potentially creating another further down the glycolytic pathway. Aldolase B is just one important enzyme in one step out of several in glycolysis, so it is important to make sure that replacing aldolase B doesn't interfere with any other important steps in this process. Knockout Drosophila glycolytic gene expression should be comparable to that seen in wild-type Drosophila given the same chemical.
I hypothesize that chemicals most similar in enzymatic function and structure to aldolase B will allow for phenotypically normal fructose metabolism knockout Drosophila.
I hypothesize that chemicals most similar in enzymatic function and structure to aldolase B will allow for phenotypically normal fructose metabolism knockout Drosophila.
Specific aim 2: To determine the importance of the C-terminus region and the low complexity region (LCR) in fructose metabolism.
Reason: Researching the importance of these aldolase B protein regions in homologs such as the fruit fly may allow us to determine the function and significance of the corresponding regions in human aldolase B.
Approach:
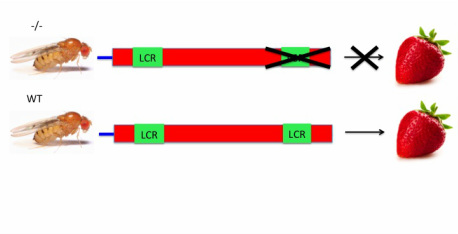
Step 1: Use RNAi to knock out the C-terminus region and (separately) the LCR in Drosophila.
Reason: Researching the importance of these aldolase B protein regions in homologs such as the fruit fly may allow us to determine the function and significance of the corresponding regions in human aldolase B.
Approach:
Step 1: Use RNAi to knock out the C-terminus region and (separately) the LCR in Drosophila.
Left: Knocking out the C-terminus region Right: Knocking out the LCR
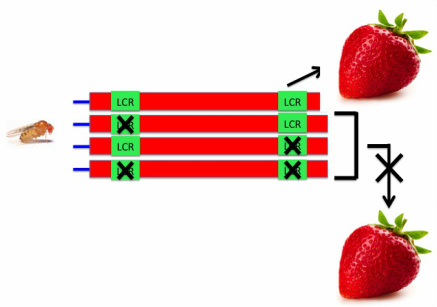
Step 2: Give Drosophila fructose and examine their phenotype. I hypothesize that knockout Drosophila for the C-terminus region will be unable to process fructose and die because the C-terminus region contains sites commonly mutated in cases of HFI. Additionally, knockout Drosophila for the LCR will be unable to process fructose and die because the LCR is important for proper enzymatic folding.
Left: Expected results for knocking out the C-terminus region Right: Expected results for knocking out the LCR
These experiments have the potential to give new insight on possible chemical supplements for aldolase B and further knowledge on the function of the C-terminus and LCR regions of aldolase B in humans. Further research idea and overall conclusions can be found here.
References
1. Cox, T.M. (1994). Aldolase B and fructose intolerance. The FASEB Journal, 8(1), 62-71.
2. James, C.L., Rellos, P., Ali, M., Heeley, A.F. & Cox, T.M. (1996). Neonatal screening for hereditary fructose intolerance: frequency of the most common mutant aldolase B allele (A149P) in the British population. Journal of Medical Genetics, 33(10), 837–841.
3. Kajihara, S., Mukait, T., Arai, Y., Owada, M., Kitagawa, T. & Hori, K. (1990). Hereditary Fructose Intolerance Caused by a Nonsense Mutation of the Aldolase
4. Understand Dairy Sensitivity: Overview. Lactaid. Accessed 13 April 2013. http://www.lactaid.com/dairy-sensitivity
5. Ekker, S.C., and Larson, J.D. (2001). Morphant technology in model development systems. Genesis 30:89–93. http://onlinelibrary.wiley.com/doi/10.1002/gene.1038/abstract
6. Miller, D, Hannon, C, and Ganetzky. (2013). A mutation in Drosophila aldolase causes temperature-sensitive paralysis, shortened lifespan, and neurodegeneration. Journal of Neurogenetics. 26 (3-4): 317-327. doi: 10.3109/01677063.2012.706346.
7. Thurston, J.H., Jones, E.M., Hauhart, R.E. (1974). Decrease and Inhibition of Liver Glycogen Phosphorylase After Fructose. Diabetes, 23 (7), 597-604. doi: 10.2337/diab.23.7.597
8. NCBI BLAST. Homo sapiens chromosome 9, GRCh37.p13 Primary Assembly compared to Mus musculus aldolase B, fructose-bisphosphate, mRNA (cDNA clone MGC:36362 IMAGE:4975382), complete cds. http://blast.ncbi.nlm.nih.gov/Blast.cgi
Drosophila image courtesy of http://obbard.bio.ed.ac.uk/photo_gallery/Drosophila_kuntzei_lab.html. Strawberry image courtesy of http://www.figtreeolivebranch.com/how-to-hull-a-strawberry. Drosophila aldolase B diagrams created by Maddy Ford, 2014.
2. James, C.L., Rellos, P., Ali, M., Heeley, A.F. & Cox, T.M. (1996). Neonatal screening for hereditary fructose intolerance: frequency of the most common mutant aldolase B allele (A149P) in the British population. Journal of Medical Genetics, 33(10), 837–841.
3. Kajihara, S., Mukait, T., Arai, Y., Owada, M., Kitagawa, T. & Hori, K. (1990). Hereditary Fructose Intolerance Caused by a Nonsense Mutation of the Aldolase
4. Understand Dairy Sensitivity: Overview. Lactaid. Accessed 13 April 2013. http://www.lactaid.com/dairy-sensitivity
5. Ekker, S.C., and Larson, J.D. (2001). Morphant technology in model development systems. Genesis 30:89–93. http://onlinelibrary.wiley.com/doi/10.1002/gene.1038/abstract
6. Miller, D, Hannon, C, and Ganetzky. (2013). A mutation in Drosophila aldolase causes temperature-sensitive paralysis, shortened lifespan, and neurodegeneration. Journal of Neurogenetics. 26 (3-4): 317-327. doi: 10.3109/01677063.2012.706346.
7. Thurston, J.H., Jones, E.M., Hauhart, R.E. (1974). Decrease and Inhibition of Liver Glycogen Phosphorylase After Fructose. Diabetes, 23 (7), 597-604. doi: 10.2337/diab.23.7.597
8. NCBI BLAST. Homo sapiens chromosome 9, GRCh37.p13 Primary Assembly compared to Mus musculus aldolase B, fructose-bisphosphate, mRNA (cDNA clone MGC:36362 IMAGE:4975382), complete cds. http://blast.ncbi.nlm.nih.gov/Blast.cgi
Drosophila image courtesy of http://obbard.bio.ed.ac.uk/photo_gallery/Drosophila_kuntzei_lab.html. Strawberry image courtesy of http://www.figtreeolivebranch.com/how-to-hull-a-strawberry. Drosophila aldolase B diagrams created by Maddy Ford, 2014.